The DNA Network |
| Researchers find way to make tumor cells easier to destroy [Think Gene] Posted: 06 May 2008 05:42 PM CDT Tumors have a unique vulnerability that can be exploited to make them more sensitive to heat and radiation, researchers at Washington University School of Medicine in St. Louis report. The Washington University radiation oncology researchers found that tumors have a built-in mechanism that protects them from heat (hyperthermia) damage and most likely decreases the benefit of [...] |
| Posted: 06 May 2008 05:41 PM CDT Analysis and diagnosis in a chip format are coming of age, but their practical application has been limited because until now, the sample usually had to be prepared separately and on a nonminiaturized scale. Jürgen Pipper and his team at the Institute of Bioengineering and Nanotechnology in Singapore want to change this. They have now [...] |
| [Confirmed] Man Bear Pig Sighting in Wellston, Ohio, Appalachia! [Think Gene] Posted: 06 May 2008 04:38 PM CDT Breaking news: Think Gene has recently received a civilian report [edit: confirmed] of a Man Bear Pig sighting in Appalachian Ohio! I would like to know the sponsoring agency for the thinkgene website. today my students were surfing the web and found your man, bear, pig [editor: Man Bear Pig, April 1st 2008]. At first glance [...] |
| Oppose Louisiana Senate Bill 733, the Louisiana Science (mis)Education Act. [Synthesis] Posted: 06 May 2008 04:36 PM CDT
.gif) This post is about creationism, education, evolution, intelligent design This post is about creationism, education, evolution, intelligent designRelated posts |
| How many genes do you REALLY have? [adaptivecomplexity's column] Posted: 06 May 2008 04:35 PM CDT You've probably heard widely varying estimates for the number of protein-coding genes in the human genome. Back before the genome sequence came out, many scientists guessed that the number was around 100,000. When scientists first looked at the newly completed human genome sequence in 2001, they found about 27,000 genes, and ever since then I have seen estimates ranging from 20,000 to 30,000. |
| DDC Announces New 3-Day Turnaround Time for DNA Paternity Tests [The DNA Testing Blog] Posted: 06 May 2008 04:32 PM CDT DNA Diagnostics Center is pleased to announce that its laboratory has shortened its turnaround time for DNA paternity tests from 5 business days to 3 business days. This shift marks a major achievement in the DNA testing industry, where a 5 working-day turnaround time is the standard for paternity tests. "We are leading the way by [...] |
| Scientists identify interacting proteins key to melanoma development, treatment [Think Gene] Posted: 06 May 2008 04:27 PM CDT Researchers have discovered how a mole develops into melanoma by showing the interaction of two key proteins involved in 60-70 percent of tumors. The Penn State scientists also demonstrate that therapeutic targeting of these proteins is necessary for drugs to effectively treat this deadly form of cancer. “We have shown that when two proteins – [...] |
| The cooperative view: New evidence suggests a symbiogenetic origin for the centrosome [Think Gene] Posted: 06 May 2008 04:26 PM CDT There are two ways in which cooperation is the theme of a paper published this week by Mark Alliegro and Mary Anne Alliegro, scientists at the Marine Biological Laboratory's (MBL) Josephine Bay Paul Center. One is revealed in the paper's acknowledgments, where the Alliegros thank those who helped them after Hurricane Katrina completely disrupted their laboratory [...] |
| Blocked brain enzyme decreases appetite and promotes weight loss [Think Gene] Posted: 06 May 2008 04:24 PM CDT Imagine being able to tone down appetite and promote weight loss, while improving the body's ability to handle blood sugar levels. That's just what Tony Means, PhD, and his team at the Duke University Medical Center were able to do when they blocked a brain enzyme, CaMKK2, in mice. "We believe we have identified an important drug [...] |
| There’s very good agreement among 23andme and decodeme genetic profiles. [Synthesis] Posted: 06 May 2008 03:44 PM CDT Antonio C B Oliveira had himself tested and wrote a little program to compare the results, finding only 23 discrepancies out of 560299 calls made by both services. Megan Smolenyak’s husband’s tests from both services were compared by Ann Turner, who found 35 disagreements among 560128 co-calls. The called disagreements are fewer than the no call differences, which certainly seems like an acceptable rate to me. I wonder what the disagreements would be if someone were to have themselves tested twice, using two independent samples, say 6 months apart? The Genetic Genealogist has a nice summary. .gif) This post is about 23andme, decodeme, personal genomics, SNP This post is about 23andme, decodeme, personal genomics, SNPRelated posts |
| A career in science is a bad decision for a smart man. [Synthesis] Posted: 06 May 2008 03:17 PM CDT Philip Greenspun, an entrepreneur who became successful at software development after completing a PhD in EE, has a popular essay on careers in science. I tend to agree with his pessimistic viewpoint, but I do think that things are a little different in medical research than physics. I think jobs are a little easier to come by, for one. I think it’s also important to keep in mind that his perspective is based on the extremely competitive east coast academic environment, and that he actually has been successful in the non-academic route. The main point he seems to be making is that there are less women than men in academic science careers, not because women are less capable or more concerned with family life or anything, but rather that there are more men simply because men are the only ones stupid enough to let their ego influence their career choice.
I know that my sense of identity and self-worth is tied very closely to my career. When things are going well, I feel like I’m doing the right thing, but when I can’t get anything to happen, I get depressed. I can certainly understand that women, who tend to experience assaults upon their self-worth from all directions from an early age, are a little better at eventually choosing more appropriate things to base their self-esteem upon. via infoproc, some commentary from the life science POV at Bayblab .gif) This post is about career, science This post is about career, scienceRelated posts |
| What can happen when geneticists play God: [adaptivecomplexity's column] Posted: 06 May 2008 12:54 PM CDT Nerdy comics don't get any funnier than xkcd: 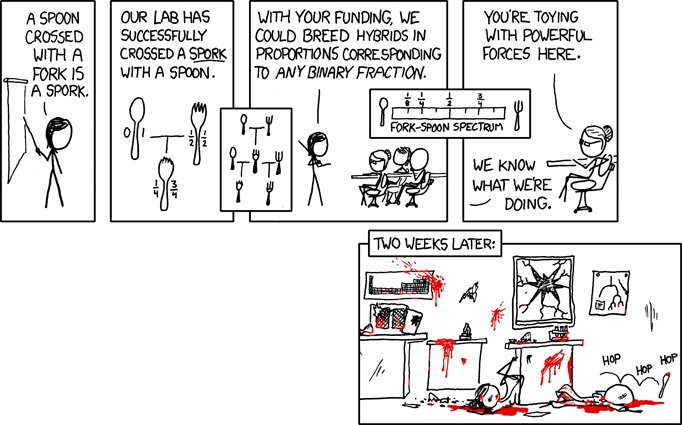 |
| Greatest application of physics and the internet EVAR! [Bayblab] Posted: 06 May 2008 12:08 PM CDT In the quest for perfect BT (bra technology), the folks at 'shock absorber' have managed to create a mesmerizing website. To check out the bounceometer be sure to have your cup size and an estimation of your level of activity ready, and watch the efficiency of the 'shock absorber'. |
| Using Disposable Income for Genetic Tests [Eye on DNA] Posted: 06 May 2008 11:29 AM CDT The New York Times reported this past weekend that more people are having problems obtaining affordable health insurance. On top of the budget constraints people face during a recession, even those who are covered by employer health insurance have to deal with “some combination of higher premiums, less extensive coverage, and bigger out-of-pocket deductibles and co-payments.” This means that many skip routine check-ups and avoid seeing the doctor unless absolutely necessary.
How does this affect the potential market for genetic services? If people can’t even afford to pay for necessary maintenance medication, eye glasses, or diabetes test strips, how do personal genomics companies expect to expand their market for elective health services? And yet, direct-to-consumer genetic testing is more widely available in the U.S. than in any other country. Where socialized medicine prevails in countries such as the UK, Singapore, and Iceland, it seems that people would have more disposable income to spend on optional healthcare. It would be interesting to see the uptake of personal genomic services in countries other than the U.S. although culture and legalities would be important factors as well. For example, are Icelanders more interested in and willing to spend money on personal genomics given that one of the more successful personal genomics companies, deCODE genetics , is based in Iceland and has published studies closely examining its citizens? Eventually, personalized medicine incorporating genetic information will become a fact of life. At that point, genetic testing will be routinely covered by insurance as with any other laboratory test or become a hidden cost when pricing pharmaceuticals, i.e., a pharmaceutical company would cover the cost of a genetic test in order to determine type and dosage of a particular medicine. For now, however, it seems that it would be hard for most people to justify spending any of their disposable income on genetic tests or scans unless family history or other known medical conditions alert them to the need for extra information and vigilance. How are you using your disposable income? |
| Lancet and the Sherpa [The Gene Sherpa: Personalized Medicine and You] Posted: 06 May 2008 10:34 AM CDT |
| Lucky frenchman walks away with MacBook Air and CLC Genomics Workbench [Next Generation Sequencing] Posted: 06 May 2008 09:30 AM CDT At the recent Bio-IT World Conference in Boston, USA, we held a competition where the lucky winner could walk away with a MacBook Air and an extensive collection of CLC bio software, including CLC Genomics Workbench. The lucky winner was Sébastien Vachenc from Laboratoires Fournier - a Solvay Pharmaceuticals company. He was of course most thrilled [...] |
| Osteoporosis and Gene Tests [The Gene Sherpa: Personalized Medicine and You] Posted: 06 May 2008 08:16 AM CDT |
| New “research” web-pages at BiRC [Mailund on the Internet] Posted: 06 May 2008 07:37 AM CDT Yesterday I updated the “Research” page at the BiRC website. We have been wanting to update the description of our research for a while. It is after all an important part of the website of a research institution, and ours was hopelessly outdated and a mess of small and large projects, some we were actively working on and some that were little more than areas we were interested in. A year ago, we had a big meeting and discussed, all of BiRC, for an entire afternoon how to update the pages. It really goes without saying that nothing could be decided in such a large group. Instead, around Christmas, Storm, our director, called a meeting inviting those interested in updating our web-pages and willing to put in the work to implement the changes. Requiring that people should back up their ideas with actual work reduced the group significantly. We were only three, myself, Enette and Storm. I got the task of coming up with a draft of the research pages and present it to the rest of BiRC. What is a research project and what goes on a research center’s homepage?The first thing I had to decide was what I wanted on the page. The form of the content is usually easier to figure out once you know what the content is. On a webpage describing the research going on at BiRC, I wanted it to be the actual research going on at BiRC. This sounds obvious, but I have seen many research institutions listing their research interests with lists I find completely unbelievable. You might be interested in a lot of different problems, but there is a limit to how many fields you can make an active contribution to. I find it dishonest if you claim to be doing research in field where, really, all you have done is publish a paper related to the field five years ago, and you occasionally read a paper about the topic. Too often, that is what I have noticed. There is nothing wrong with focusing on a few research areas at a research institution. Sure, it is fun to do a few projects outside your main research area from time to time, but if people read the web pages to find out what you are doing at the institution, then they should find the area where you are spending the majority of your time, not something you think about every second year. If a post doc wants to come and work at BiRC, he should know what kind of work we actually do, not what kind of work we like to read about. Anyway, this is turning into a bit of a rant, but that is the kind of thoughts I was having at the time. So, I wanted the page to contain our active research areas, and I wanted to back up the “we are actively working on this stuff” claim we are implicitly making when describing research on our web pages. How do you prove that you are active? You put up the money or the work! Ideally, you want to prove that you are doing research in an area, by showing that you are actively publishing in it. If you do not publish, are you really doing research there? If you are, is it any good? I wanted each project to be able to show at least two-three papers a year to be considered active. I made an exception if there was funding for a project, but it hadn’t really produced any results yet… I am still not sure this is a good idea, but my arm was twisted a bit to allow for this… Designing the web pagesAfter these considerations, I started to design the pages. I made an overview page, where people can scan through all the research areas at BiRC very quickly, and with links to sub-pages — one for each research area — that can contain more information. Putting it togetherThis design I presented to BiRC and no one objected. Good. Then I asked people to send me the projects (description and publication lists) they wanted on the page. Deadline March 15, which would give me time to set up the sub-pages and such, and then the new page could be put on the on May 1. Naturally, but March 16 I hadn’t received anything and by May 1 I had a few half-baked descriptions. Deadlines are rarely taken that serious when they are internal deadlines like this. I am pretty bad at meeting them myself, so I cannot really complain. With a lot of threats I got the material I needed over the weekend — well, enough to make descriptions of the main research areas at BiRC — and I put the pages together. I’m personally maintaining more of them than I had planned. I really only wanted to be responsible for the Association Mapping page. This is my own main area, and the one where I consider myself the local expert. I ended up maintaining virtually all the pages where I am contributing anything. I plan to push the responsibility to someone else when I get the chance, but for now this is how it is. |
| On the road again… [Mailund on the Internet] Posted: 06 May 2008 05:33 AM CDT I’m in London now, working with David Balding on HapCluster. We had a late dinner yesterday and I am slightly hung over and a bit lacking in sleep, which is not the optimal state of mind to be trying to solve a mixing problem in an MCMC… |
| Humanity’s Brush With Extinction [] Posted: 06 May 2008 02:16 AM CDT
Turns out our DNA is keeping better records than historians. Among the many interesting historical events DNA reveals about humans is a reduction of the worldwide human population to about 2,000 individuals around 70,000 years ago.
The findings come from the Genographic Project, launched in 2005 to study anthropology using gene sequencing and bioinformatics. The DNA record coincides with geological records strongly suggesting a correlation with extreme drought during the era of near-extinction. Genographic Project Paleontologist Meave Leakey: “Who would have thought that as recently as 70,000 years ago, extremes of climate had reduced our population to such small numbers that we were on the very edge of extinction?” |
| Would you Sterilise Growth Media With A Microwave? [Bitesize Bio] Posted: 06 May 2008 01:00 AM CDT We have had a rush on time and money saving techniques on Bitesize Bio in the last few weeks. Ways to re-cycle electroporation cuvettes, reduce gel buffer costs, do fast restriction digests and re-cycle midiprep columns have all been suggested. In this article I’ll add the possibility of using a microwave to sterilize or decontaminate growth media. From the outset I’d like to say that I am not too sure about this, but I’ll make the case and you can tell me what you think. Normally, growth medium is sterilized or decontaminated using an autoclave. Autoclaves are generally expensive, energy-hungry beasts that (in my experience) break down a lot so I would be very happy to use them less if I could. The case for using microwave ovens for decontamination of cultures or materials was made back in 1977 by Latimer and Matsen. They showed that 1-5 minutes in a conventional microwave was sufficient to decontaminate 5mL cultures or petri dishes of common clinical pathogens including E. coli, S. aureus and K. pneumoniae. B. subtilis spores proved a bit more subborn, requiring more than 10 minutes of microwaves to wipe them out. Border and Rice-Spearman backed this up with a 1999 study that showed materials contaminated with various bacteria and yeast strains were completely decontaminated by one minute in the microwave (I guess their microwave was better). And in 2006, Silva et al, investigating the decontamination of dentures, showed that 6 minutes in the microwave sterilised S. aureus and C. albicans but only partially disinfected P. aeruginosa and B. subtilis. Sterilisation using microwaves A 2001 Biotechniques paper by Weiss and Galande showed that LB plates made from microwave-sterilised LB-agar were apparently sterile (control plates were no detectably contaminated by microorganisms), had a similar shelf-life to autoclaved plates and supported bacterial growth as normal. The plates were prepared from dry powders dissolved in distilled water and aliquoted into 50mL tubes. This is a very fast way to make plates and has the added advantage that the antibiotics can be added in from the start as they are not destroyed by microwaves. Invitrogen have a product that takes advantage of this. ImMedia is LB medium provided as sachets of dried, weighed power containing all of the required media components (including antibiotics). It is designed so that you can just add the sachet contents to water, microwave and your media is ready. But Weiss and Galande’s method is just as good and much cheaper. My view My take-home from this is that microwaves are are reasonably good at decontamination but more stubborn microorganisms (e.g. the spore-forming B subtilis) are not effectively disinfected. So the method does not sound too reliable to me. Also, filling the lab with smelly fumes from contaminated stocks does not seem to be a good idea. For easy liquid culture decontamination I think I will stick with Virkon, and for solid media, autoclaving seems to be the only good option. The microwave media prep method is certainly interesting. Weiss and Galande’s results seem to be pretty robust and I would consider this method for an emergency media prep - if I need to start an E. coli culture last thing at night and there’s no sterile media available. Although the decontamination results show that microwaves don’t kill everything, E. coli grows so quickly that for routine purposes, a low level of contamination by slower growing organisms can be tolerated. But I would not use this method for anything other than routine cultures and certainly not for slow growing organisms. Maybe that’s just me being a typical scientist, reluctant to take on new methods as Liam suggested. The microwave method is also limited by the fact that only small volumes (50ml) can be sterilised so it is never going to replace the autoclave for batch media production. That’s my view - what’s yours? |
| HEALTH Highlights - May 6th, 2008 [Highlight HEALTH] Posted: 05 May 2008 11:25 PM CDT
.gif) Thank you for subscribing by RSS or email. I work hard to make the articles on Highlight HEALTH engaging and I truly appreciate your interest and readership! Thank you for subscribing by RSS or email. I work hard to make the articles on Highlight HEALTH engaging and I truly appreciate your interest and readership!This article was published on Highlight HEALTH. Related articles |
| Posted: 05 May 2008 11:10 PM CDT The model fungus Podospora anserina (P. anserina) has undergone substantial evolution since its separation from Neurospora crassa, as revealed from the Podospora draft genome sequence published in BioMed Central's open access journal, Genome Biology. The study also shows that the Podospora genome contains a large, highly specialised set of genes potentially involved in the breakdown [...] |
| BILBO1 a bearer of bad fortune for trypanosomes [Think Gene] Posted: 05 May 2008 11:08 PM CDT Trypanosoma brucei is a pathogen that causes epidemics of human and animal sleeping sickness in central and Southern Africa, a disease that is fatal if untreated. A new paper published this week in the open-access journal PLoS Biology investigates the formation of a trypanosome structure called the Flagellar Pocket (FP). The work, by Derrick Robinson [...] |
| Specific gene increases susceptibility to breast cancer [Think Gene] Posted: 05 May 2008 11:06 PM CDT Much work has been done to identify genetic variations that predispose women to breast cancer. Previous work showed that variants in the gene called fibroblast growth factor receptor 2 (FGFR2) were associated with increased risk of the disease, but how these variants translated into increased risk was unknown. A new paper by Kerstin Meyer and [...] |
| Immune exhaustion in HIV infection [Think Gene] Posted: 05 May 2008 11:04 PM CDT It's the virus, stupid: immune exhaustion in HIV infection As HIV disease progresses in a person infected with the HIV virus, a group of cells in the immune system, the CD8+ T lymphocytes, become "exhausted," losing many of their abilities to kill other cells infected by the virus. For many years scientists have debated whether this [...] |
| You are subscribed to email updates from The DNA Network To stop receiving these emails, you may unsubscribe now. | Email Delivery powered by FeedBurner |
| Inbox too full? | |
| If you prefer to unsubscribe via postal mail, write to: The DNA Network, c/o FeedBurner, 20 W Kinzie, 9th Floor, Chicago IL USA 60610 | |



No comments:
Post a Comment