The DNA Network |
| A society where ideology is a substitute for evidence can go badly awry [Tomorrow's Table] Posted: 13 Aug 2008 08:14 PM CDT Olivia Knudson has a lovely piece today on why it is important for evolution to be taught in beginning biology classes. One important point that she makes is this: "A society where ideology is a substitute for evidence can go badly awry." She goes on to say: "In his book, 'The Republican War on Science,' the journalist Chris Mooney argues persuasively that a contempt for scientific evidence — or indeed, evidence of any kind — has permeated the Bush administration's policies, from climate change to sex education, from drilling for oil to the war in Iraq. A dismissal of evolution is an integral part of this general attitude". I would argue (as does she) that we see this substitute for ideology over science on the left end of the political spectrum as well. In the case of genetically engineered crops, for example, there is a pervasive ideology that GE crops are harmful to human health and to the environment, even though there has not been a single case of harm in over 10 years of cultivation. |
| TravelBlogue, or How to live vicariously through one's student. [Genomicron] Posted: 13 Aug 2008 06:08 PM CDT The very first post here was called "My grad student made me do it", and explained that a then-newly-arrived PhD student in my lab was a blogger and got me interested in blogging. He is still a blog author, and most recently has posted a very enjoyable series about his travels from more or less the bottom to the top of the USA/Canada parts of North America looking for aquatic creatures. I personally did not get to go anyplace exciting this summer, but it has been great having the option to live vicariously, especially as he was most recently at one of the coolest (literally?) places on Earth: Devon Island in the high Arctic. Follow his adventures:  Florida to Guelph: 
Guelph to Thompson: 
Churchill: 
Resolute and True Love:
|
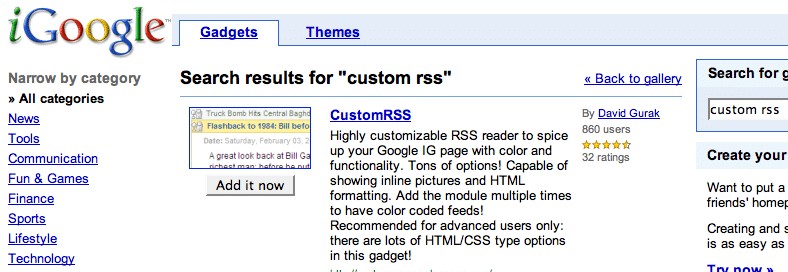
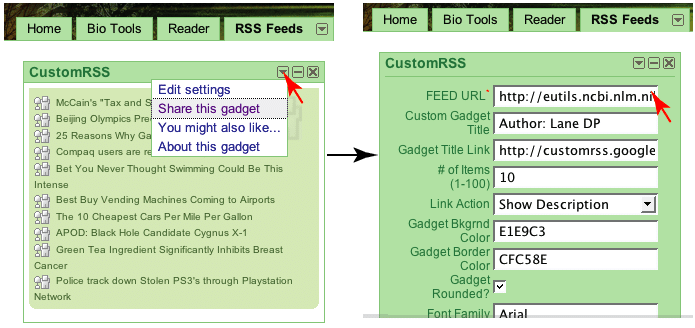
| Mendeley goes public and gets some love from ex-Skype and last.fm [My Biotech Life] Posted: 13 Aug 2008 04:18 PM CDT
Since then, Mendeley has moved into public beta, therefore anyone can register and download the software. Another great development is that the software that was originally only available for Windows users (like myself!), is now available in Mac and Linux flavors too. A press release was sent out this week where some of the project funding info was provided and I’m glad to see some respectable online company names supporting this great project, like former Skype engineers and last.fm Chairman too. If you didn’t get a code to try out Mendeley during the private beta, go give it a try now. I really recommend it. Mendeley goes public and gets some love from ex-Skype and last.fm |
| You could win an iPod and a MacBook Air and an Apple TV [Discovering Biology in a Digital World] Posted: 13 Aug 2008 01:37 PM CDT if you take this survey. Wanna change the world? Make it possible for everyone to talk about science in a normal conversation? Do you have ideas for improving science literacy? Seed is interested in your ideas. Answer the survey and share your thoughts. And I've seen the MacBook Air. It's beautiful. UPDATE: if you had trouble accessing the survey, try it again. It will be open until Friday, August 15th, 11pm EST. Read the comments on this post... |
| Benevolent dictators have longer AVPR1a promoters [biomarker-driven mental health 2.0] Posted: 13 Aug 2008 12:56 PM CDT The small neuropeptides oxytocin (OT) and arginine-vasopressin (AVP) are well known for their influence on promoting warm-and-fuzzy social behaviors in mammals. The G-protein coupled OTR and AVPR1a receptors are also the subject of much research in this area - particularly AVPR1a - since it shows differences in brain expression in polygamous vs. monogamous vole species, and also shows genetic associations with dysfunction in human social affiliation. In a recent foray into this line of research, Richard Ebstein and colleagues examine whether an individual’s willingness to give away a cache of money is related to genetic variation in the promoter of the AVPR1a. In their paper, “Individual differences in allocation of funds in the dictator game associated with length of the arginine vasopressin 1a receptor RS3 promoter region and correlation between RS3 length and hippocampal mRNA“, the researchers asked 203 college students to play the “dictator game” where, simply, one person gets a sum of money and can choose to keep it or give some of it away to the other player. Thats it. Give some of it away if you like, or just walk away with all of it, no questions asked & no consequences (your identity and the identity of the other player are masked). Amazingly, individuals actually DO give some of the money away (15% gave none of it away, 35% gave half away and 7% gave all - yes, all of it - away) … and more amazing still … those with longer stretches of microsatellite repeats at the RS1 & RS3 promoter sites in the AVPR1a gene, gave away significantly more money than individuals with shorter version of the repeats. |
| Damn slow webpages [Mailund on the Internet] Posted: 13 Aug 2008 10:15 AM CDT If you go to BiRC’s homepage, and I suggest you don’t right now, you will experience an extremely slow load time. If you are lucky, your browser will crash. If you are really lucky, your computer will crash. We have this problem, you see. The CSS file is 31Mb and the Javascript file is 28Mb. It seems that every time the page has a hit, a few lines are added to each of these files. Why this happens has been bugging us a little while. We are using a CMS developed in-house. It’s called Skeletonz and is developed by Amir Salihefendic who was employed as student programmer at BiRC a few years back, where he developed Skeletonz. Generally, it is working fine and we are very satisfied with it, but it is beta and since Amir left BiRC we haven’t had much support on it. Anyway, back to the mystery of the growing files… We have suspected the blog plugin for a while. The lines added to the files concerns the blog, so that makes sense. However, the problem didn’t go away when I disabled the blog, so I didn’t think it was the blog anyway… Now I think it is again. I think it was enough to install the blog plugin. I’ve found a bit of suspicious code I think is causing the problem. I do not have the access rights to change the code, but I’ll get that and try my fix tomorrow or next week. It is a bit embarrassing that an informatics group has been unable to have a homepage up and running without problems, but there you go. I just hope we’ve solved the problem now… |
| our new pub @ nrn - yay! [biomarker-driven mental health 2.0] Posted: 13 Aug 2008 09:43 AM CDT |
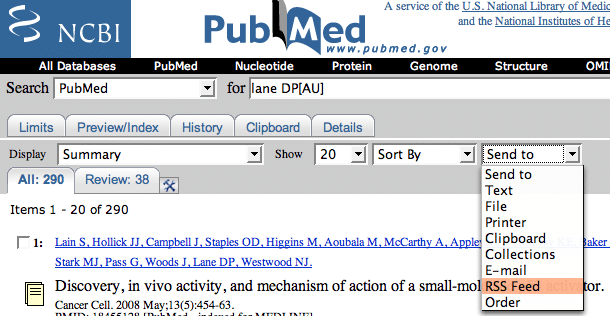
| Pubmed + RSS + iGoogle = Easy Lit Updates [Bitesize Bio] Posted: 12 Aug 2008 11:59 PM CDT We’ve talked before about ways to use technology to help you with the vital job of keeping up with the literature (see here and here). Now here is another one to add to the list. This approach was first flagged up by Eric (thanks Eric!) in a comment on Carrie’s article about improving your Pubmed searches. The idea is to use the combined power of Pubmed, RSS feeds and iGoogle to create a page of RSS feed boxes that will keep you continually updated on articles containing your keywords of interest, or from specific authors or journals. It is nice and simple, but I find it an incredibly powerful and fast method of literature scanning compared to email updates or browsing each journal individually. So here’s how to set it up. 1. If you haven’t already got one, you’ll need an iGoogle account. Just sign up at iGoogle.com. 2. In your iGoogle account, click the “add a tab” button to add a new tab, and name it “RSS Feeds”, or whatever you like. 3. Now go to Pubmed and search for a keyword, author or journal (or any combination of these you want) on which you want to be kept updated. Read Carrie’s article on Pubmed searches for some great ideas on how to perform optimal searches. One especially good tip is to use field tags to target your search: e.g.
4. When you get the search results, click on the “send to” drop-down menu and choose “RSS” feed.
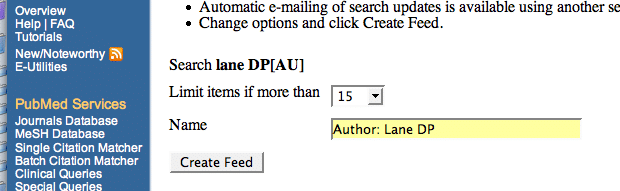
5. In the window that you are sent to, choose a title for the feed and the number of articles you want to be displayed. Then click “create feed”. 6. Clicking on the orange XML button will then take you to your feed. Copy the feed address (URL) from your browser’s address window. 7. Now you need to add a feed reader to your iGoogle page. iGoogle has many available RSS feed gadgets but the best are: Simple rss reader, CustomRSS, Slim RSS Reader, and Feeds in Tabs. To add one of these click on the “Add stuff” link then search for the gadget (or just search for “RSS reader” and choose your own) and click “Add It Now” to add it to your iGoogle page 8. Back in your newly created iGoogle RSS feed page, click the down arrow>edit settings in the RSS feed box you have justed added. Paste your feed address into the “FEED URL” box and add the title of your choice to the “Custom Gadget Title” box, then hit “save”. 9. To add another feed, go back to step 3. Repeat until you have a page full of feeds than you can easily scan to keep up with your various literature focuses. Remember that in iGoogle you can drag and drop your various feed gadgets so that they are ordered on the page in the way that you want. Think this is any good? Got a better way to keep up with the literature? Let us know in the comments… |
| The Promise of Stem Cells to Repair the Heart [Highlight HEALTH] Posted: 12 Aug 2008 11:32 PM CDT
Animal embryos differentiate into three cellular layers called germ layers: the ectoderm, mesoderm and endoderm. Tissues and organs of the body develop from these germ layers through further differentiation. The mesoderm germ layer differentiates into circulatory and urogenital systems, connective tissue, muscle and bone. Embryonic stem cells have the potential to develop into almost any type of cell in the body. To be used therapeutically, however, scientists have to understand how to direct stem cells to become specialized cell types, such as skin or heart cells. Why use stem cells to repair the heart? Heart muscle cells from a patient generally won’t divide in a sufficient number to replace damaged areas. Taking a part of the heart muscle from another area only creates more damage. Embryonic stem cells thus provide an external and abundant source of cells for heart muscle repair. Scientists at Washington University School of Medicine in St. Louis found that expression of the gene Mesp1 induced expression of mesodermal markers and genetic changes associated with the transition from embryonic to epithelial tissue (termed epithelial-mesenchymal transition). A typical epithelium is a sheet of cells, often one cell thick, held tightly together by cell-cell junctions in a uniform array. Adhesions between neighboring epithelial cells inhibit movement of individual cells. In contrast, mesenchymal cells have neither organized structure nor tight intracellular adhesion, allowing for increased migratory capacity. Cells undergoing the epithelial-mesenchymal transition (EMT) experience transient structural changes that result in a loss of contact with neighboring cells and a gain in motility. This process is vital to movements that reorganize the embryonic germ layers and to the development of other migratory cell types [2]. Many of the changes associated with cells undergoing developmental EMTs are also observed in wound healing, fibrosis and cancer. Using mouse embryonic stem cells, researchers showed that Mesp1 expression restricted the potential fates, generating progenitors or precursor cells with the potential to differentiate into cardiovascular cells but, importantly, not into hematopoietic cells (meaning blood-forming cells). They further demonstrated that Mesp1 induces expression of genes specific to cardiovascular development. The authors suggest that Mesp1 may selectively program the development of endothelial, cardiac and smooth muscle cells. According to senior author Kenneth Murphy, M.D., Ph.D., senior author and Professor of Pathology and Immunology at Washington University School of Medicine [3]:
This work has the potential to one day treat cardiovascular diseases using human stem cells. Scientists next plan to identify gene programs and map out the pathways that specify development of the three cardiac cell types: endothelial, cardiac and smooth muscle cells. For more information on stem cells and the repair of a damaged heart, see Stem Cell Information at the NIH. References
.gif) Thank you for subscribing by RSS or email. I work hard to make the articles on Highlight HEALTH engaging and I truly appreciate your interest and readership! Thank you for subscribing by RSS or email. I work hard to make the articles on Highlight HEALTH engaging and I truly appreciate your interest and readership!This article was published on Highlight HEALTH. Other Articles You May Like |
| Basic concepts in science: Crop genetic engineering [Tomorrow's Table] Posted: 12 Aug 2008 10:03 PM CDT Some scientists and policy decision-makers have proposed that genetic engineering (GE), a modern form of crop modification, will help create a new generation of plants that will dramatically reduce our dependence on pesticides, enhance the health of our agricultural systems, and increase the nutritional content of food. They believe GE will be a dramatic step forward that will topple decades of criticism about the dangerous overuse of pesticides and toxic herbicides, leading us to a more ecological way of farming. Or will it? While the public has generally accepted the application of GE for the production of new medicines, some consumers indicate grave unease over the consumption and production of GE food, viewing it as unnatural, potentially unsafe to eat and environmentally disruptive. Of these skeptics, the organic farming community has been particularly vocal in its criticism. Some consumers believe that because organic farmers have learned how to produce healthy nutritious food, GE plants are not needed. Definition: GE is not a farming method. It is a modern form of crop modification that differs from plant breeding in two basic ways: 1. Plant breeding allows gene transfer only between closely related species. With genetic engineering, genes from the same species or from any other species, even those from animals, can be introduced into a plant. Therefore genetic engineering creates a vast potential for crop alteration. 2. Plant breeding mixes large sets of genes of unknown function, whereas genetic engineering generally introduces only one to a few well-characterized genes at a time. History: For thousands of years, farmers have deliberately selected and improved plants with desired characteristics from wild and cultivated plants. For example, 10,000 years ago farmers in ancient Mesopotamia developed a hybrid between wild species of wheat and cultivated wheat that became the ancestor of our modern bread wheat. Today breeders manipulate plant species to create desired combinations of traits for specific purposes. In rice and corn, this artificial selection process is generally carried out using pollination. In this process the breeder transfers the male pollen grains to the female part of the flower. Using these techniques, breeders have been highly successful in developing new varieties; so successful in fact that corn varieties only faintly resemble their predecessors and survive only in human-made environments. Today, all of our principal food crops come from domesticated varieties. As with breeding, the goal of genetic engineering is to alter the genetic makeup of the crop. The Process of plant genetic engineering There are a couple of ways to genetically engineer a plant. One method is to use particle bombardment, or biolistics. In this process, millions of DNA-coated metal particles are shot at target cells or tissues using a biolistic device or gene gun. The DNA elutes off the particles that lodge inside the cells, and a portion may be stably incorporated in the host chromosomes. Another method is called "Agrobacterium-mediated" transformation. For rice, the starting material is an immature and still doughy seed, which carries the precious embryo and the genetic material within. If left to mature, the embryo would become edible grain. A quick dip in ethanol, followed by a soak in bleach and a spray of sterile water will protect the rice embryo from contaminants in the air—bacteria and fungi that are harmless to humans but can kill the delicate rice cells. The hulled grains are placed onto a freshly prepared plate carrying nutrients that will nourish the embryo and moved to a growth chamber that will supply adequate light and heat. In about two weeks, the grain will grow into a glistening mass called a "callus," a sort of stem cell open to direction, not yet having decided whether to develop into a particular organ, such as a leaf or root. The new callus is separated from the grain with tweezers. The next step is to introduce a new gene into the cells of the rice callus; to do this scientists rely on a soil bacterium called Agrobacterium. This bacterium can do something that virtually no other organism can do; it can form a bridge to the plant cell and then transfer some of its own genes across the plant cell wall, then across the membrane and into the nucleus. This ancient process, known to biologists for a century, was only understood in detail over the last thirty years. It is now known that this gene transfer "transforms" the plants into food production units for the bacteria. You can tell if a plant is infected by the appearance of large tumors at the base of the plant. Sometimes the results are dramatic—one of the oak trees on the UC Davis campus has a crown gall tumor the size of a small car. Biologists, faithful to their long tradition of manipulation and exploitation, have cleverly removed the bacterial genes that cause the tumor so that the bacteria can infect without disrupting plant growth. They have also figured out that it is possible to replace some of the bacterial genes with genes from other species. The bacteria, unaware that the genes have been swapped, will deliver the new genes into the plant. To carry out this subterfuge, biologists employ other tools of our trade: restriction enzymes that act like tiny scissors to cut out the bacterial genes and ligases that act like glue to insert the new genes into the genome of Agrobacterium. This cutting and pasting method is used to introduce genes-of-interest, say a resistance gene or stress tolerance gene into Agrobacterium. The callus can then be dipped into a broth containing the engineered bacteria. At this point, the bacterium acts like a courier delivering the gene-of-interest into the genome of the cultivated rice species. The bacteria must first identify its target, then infect the cell, transfer its DNA across the rice cell membrane into the nucleus, and finally into DNA that is bundled into chromosomes in each nucleus. In nature and in the lab, the bacteria do the work of gene delivery. Scientists cannot predict where the new gene will land beforehand, although it is straightforward to determine the location of the new gene after it is integrated into the crop DNA. Sometimes the gene will land in a spot that disrupts a critical function of the rice cell, in which case that cell will no longer grow. In most cases, however, the transformed cell will thrive and reproduce carrying a new bit of genetic material along with it. How unnatural is this? If we insert genes into random sites in a genome, won't we destabilize a structure that has evolved over millions of years? It turns out that plant genomes are used to this kind of abuse. It is now well documented that rice and other organisms contain pieces of DNA that move around (called transposable elements) in a seemingly erratic fashion. Not only do they insert themselves into new places, but sometimes they pick up pieces of other genes and take those fragments along for the ride as well. For example, a recent study showed that in the rice genome there are over 3000 of these pieces of DNA-containing fragments, called pack mules, from more than 1000 genes (Jiang et al. 2004). Sometimes several fragments are picked up from different genes, rearranged, fused, and then expressed as new proteins. By looking at the genome sequences, we also now know that plants have acquired genes from many different organisms. It seems then that the genetic engineering process we carry out in the lab has certain similarities to that which occurs in nature. Not all of the bacteria will be successful in transferring their DNA to the cell; only a very few of the rice cells will receive the new gene and be "transformed." How then does the biologist distinguish the genetically engineered cell from the thousands of rice cells lacking the genes? In fact, this would not be possible without another tool, called the marker gene. In early experiments of plant transformation, the commonly used marker was a gene that encoded resistance to an antibiotic. In this process, not only is the new gene transferred to the cell, but the marker gene is as well. Today, other markers are available, such as those that allow the transformed plant to grow on high levels of particular sugars; in essence this marker is a sugar enablement gene that allows it to grow on sugars it cannot otherwise use. By placing the infected rice cells on the "selection" (sugar or antibiotic) media only the transformed cells will survive, inhibiting growth of the rest of the plant cells. In other words, the marker genes bestow properties of survival only to those cells that are genetically engineered, allowing biologists to pluck the newly transformed cells from a lawn of dying, untransformed cells. In two more weeks the newly transformed cells will give rise to new cells carrying the marker gene and the gene-of-interest. These new cells appear as tiny clumps of whitish, nearly translucent globs about the size of small beads. These cells are genetically identical to the starting material (the rice seed), except that they also possess the desired gene-of-interest and the marker gene. Once transferred to nutrient plates containing plant hormones, the cells will produce roots and shoots. The GE seedlings can be transplanted into soil-filled pots. They will reproduce and produce seed like any other rice plant. Today, one billion acres of GE crops have been grown; hundreds of millions of people have eaten GE food for more than a decade without a single verifiable case of adverse side effects to the environment or to human health. Still, GE provokes controversy, and, sometimes, violent protests. Potential for using GE to boost crop yields One way to boost yields is to develop crops that can survive harsh conditions such as drought, cold, heat, salt, and flooding. Many of the world's poorest people farm in areas that are far from ideal, and freshwater sources are decreasing in quantity and quality throughout the world. New crop varieties designed to survive in difficult environments will have a significant human and ecological impact. In the future this is where genetic engineering will likely play an important role. Crops with enhanced tolerance to drought, for instance, would allow farmers to produce more food using less water. Already there are varieties of genetically engineered wheat that can tolerate drought, as well as rice that can tolerate flooding and tomato plants that can tolerate salt. Another important challenge is to fight pests and disease, which take an estimated 20 to 40 percent bite out of agricultural productivity worldwide. Reducing this loss would be equivalent to creating more land and more water. But current pesticide use is a health and environmental hazard, and organic and genetic engineering offer complementary solutions. Genetic engineering can be used to develop seeds with enhanced resistance to pests and pathogens; organic farming can manage the overall spectrum of pests more effectively. Genetically engineered crops have already enjoyed major success against pests. For example, on farm field trials carried out in central and southern India, where small-scale farmers typically suffer large losses because of pests, average yields of genetically engineered crops exceeded those of conventional crops by 80 percent. In Hawaii, the 1998 introduction of an engineered papaya plant that could resist the papaya ringspot virus virtually saved the industry. There was no organic approach available then to protect the papaya from this devastating disease, nor is there now. Genetic engineering can also help achieve other goals of the ecological farming movement. By reducing the use of pesticides and by reducing pests and disease, it can make farming more affordable and thus keep family farmers in business and assure local food security. It can also make food more nutritious: In 2011, plant breeders expect to release "golden rice," a genetically engineered variety that will help fight Vitamin A deficiency in the developing world, a disease that contributes to the deaths of hundreds of thousands of young children each year. Opposition to GE crops To many people, genetically altering crops feels fundamentally wrong or unnatural. They believe that farmers already have enough tools for a productive and healthy farming system. On an environmental level, many worry that genetically engineered crops will cross-pollinate nearby species to create a new kind of weed that could invade pristine ecosystems and destroy native plant populations. On a personal level, many consumers worry that genetically engineered foods are unsafe or unhealthy to eat. So far, however, it appears those concerns are driven more by technological anxiety than by science. Virtually all scientific panels that have studied this matter have concluded that pollen drift from genetically engineered varieties currently grown in the United States does not pose a risk of invasiveness. (Although this does not mean that future crop varieties will also be harmless: each new crop variety must be considered on a case-by-case basis.) And in terms of food safety, a report by the National Academy of Sciences concluded that the process of adding genes to our food by genetic engineering is no riskier than mixing genes by conventional plant breeding. Today 70 percent of all processed foods in the United States have at least one ingredient from genetically engineered corn, cotton, canola, or soybean. Unlike the well-documented adverse effects of some pesticides, there has not been a single case of illness associated with these crops. Intellectual property Many opponents of genetic engineering fear that a blizzard of patents on genetically engineered plants and seeds will put control of agriculture in the hands of a few giant companies that produce the seeds. Yet there are many new and imaginative methods that businesses and universities are now using to ensure that breakthroughs and useful technologies benefit less developed countries and small-acreage farmers. For example, the nonprofit initiative Public Intellectual Property Resource for Agriculture brings together intellectual property from more than 40 universities, public agencies, and nonprofit institutes and makes these technologies available to developing countries around the world for humanitarian purposes. |
| Tomorrow's Table in the classroom [Tomorrow's Table] Posted: 12 Aug 2008 10:02 PM CDT "I really enjoyed the book. It did a great job of keeping everything in perspective. Use again !" "Use again! A great resource and easy to understand" "The textbook was great. It had a story line to it. It was easy to remember." These are some of the comments from Oregon State University students who read the book, "Tomorrow's Table: Organic Farming, Genetics and the Future of Food". Steven Strauss, Distinguished Professor of Forest Biotechnology at Oregon State University, who directs the OSU Program for Outreach in Resource Biotechnology, chose the book for his course, which give students and the public scientifically reliable information about the use of genes and chemicals in agriculture and natural resources. Thanks Steve, for being the first to use it in the classroom! |
| U. of California can reject creationist college prep classes [adaptivecomplexity's column] Posted: 12 Aug 2008 09:46 PM CDT Religious conservatives who decry postmodernism in the academy, and object to what they see as identity-based scholarship, such a Marxist school of history or African American Studies, have no problem trying to push their own Christianity-based science on academic institutions. |
| Yeast Cells on Psychoactive Drugs [adaptivecomplexity's column] Posted: 11 Aug 2008 10:33 PM CDT When you put yeast on Prozac, do you observe any side effects? Off-target effects of psychiatric drugs are a big problem, and researchers have been eager to identify the cellular processes that are responsible for these side effects. Often the cellular process involved in the drug's the main effect is known, but what causes the side effects is not well known. To get at this issue, a group of researchers at the University of Toronto tested the effects of 214 psychoactive drugs on yeast. Why yeast? When it comes to the most basic cellular processes, yeast have the same ones we do, and in yeast it is easier to identify what parts of the cellular machinery are being affected by the drug. Even better, yeast don't have any of the sophisticated neurological machinery we have. That neurological machinery is the main target of these drugs; since that main target is absent in yeast, scientists can focus purely on the off-target drug effects. |
| You are subscribed to email updates from The DNA Network To stop receiving these emails, you may unsubscribe now. | Email Delivery powered by FeedBurner |
| Inbox too full? | |
| If you prefer to unsubscribe via postal mail, write to: The DNA Network, c/o FeedBurner, 20 W Kinzie, 9th Floor, Chicago IL USA 60610 | |




No comments:
Post a Comment