The DNA Network |
| This is Where the End of Cancer Begins - Stand Up To Cancer [Highlight HEALTH] Posted: 05 Sep 2008 07:33 PM CDT
We dreamed we’d split the atom, walk on the moon, unlock our very genetic code. To those who said ‘impossible’, ‘can’t happen’, ‘won’t happen’, we didn’t hear a word. We stood up and we changed the world. Stand up, Stand up for everyone who can’t rise anymore. One person every minute, one life in a moment is taken by a disease that we can actually cure. And in the time it’s taken to say these words, one more American has died. Unforgivable. This is where the end of cancer begins. When together we rise as one. When we Stand Up To Cancer. References
This article was published on Highlight HEALTH. Other Articles You May Like |
| Another medical chumby ... cool [biomarker-driven mental health 2.0] Posted: 05 Sep 2008 06:23 PM CDT Just saw this on engadget ... fun and useful - just like chumby but with a medical twist. Who knows, it may someday make housecalls (see link below). |
| Fat gene demonstrates role in cognitive development [biomarker-driven mental health 2.0] Posted: 05 Sep 2008 06:21 PM CDT  Image via Wikipedia Sometimes we humans tend to think we're pretty sophisticated, but let's face it, once we've got a fridge full of food and a partner to mate with, most of us - like every other species - are pretty content. So it may seem reasonable, from an evolutionary standpoint, that a gene that regulates food intake and metabolism - leptin - would have wide-ranging effects on almost every physiological system in the human body including: immune, reproduction, endocrine, skeletal and CNS. A new PLoS ONE paper entitled, "Leptin Replacement Improves Cognitive Development" reports that administration of recombinant leptin to a 5-year-old boy with a nonconservative missense leptin gene mutation (Cys-to-Thr in codon 105) yields dramatic improvements in neurocognitive function. The open access paper describes the many known effects on leptin on neuronal plasticity and it is wonderful indeed to see its success when used as a therapeutic agent. That the development of so-called 'higher' cognitive function in humans is regulated by a small peptide secreted by fat cells may be an affront to some, but not me. "Honey, pass the chicken wings !" Image via Wikipedia Sometimes we humans tend to think we're pretty sophisticated, but let's face it, once we've got a fridge full of food and a partner to mate with, most of us - like every other species - are pretty content. So it may seem reasonable, from an evolutionary standpoint, that a gene that regulates food intake and metabolism - leptin - would have wide-ranging effects on almost every physiological system in the human body including: immune, reproduction, endocrine, skeletal and CNS. A new PLoS ONE paper entitled, "Leptin Replacement Improves Cognitive Development" reports that administration of recombinant leptin to a 5-year-old boy with a nonconservative missense leptin gene mutation (Cys-to-Thr in codon 105) yields dramatic improvements in neurocognitive function. The open access paper describes the many known effects on leptin on neuronal plasticity and it is wonderful indeed to see its success when used as a therapeutic agent. That the development of so-called 'higher' cognitive function in humans is regulated by a small peptide secreted by fat cells may be an affront to some, but not me. "Honey, pass the chicken wings !" |
| Epigenetic keys to brain repair [biomarker-driven mental health 2.0] Posted: 05 Sep 2008 06:19 PM CDT  Image via Wikipedia Siming Shen et al., in their paper, "Age-dependent epigenetic control of differentiation inhibitors is critical for remyelination efficiency" provide insight on basic mechanisms of myelination. While myelination (think of it as the plastic insulation on copper electrical wires) makes normally developing neural networks much more efficient, it has a way of inhibiting the re-development and repair of mature neural circuits. The research team shows that recruitment of histone deacetylases (HDACs) is rather inefficient in mature oligodendrocytes precursor cells (the cells that adhere to bare neuronal axons and form the insulating myelin-rich sheath) in contrast to younger cells which differentiate readily. HDAC1 and HDAC2 are shown to down-regulate of Hes5 and Sox2, which have previously been implicated in blocking the differentiation of stem cells to oligodendrocytes. Here, the term 'epigenetic' refers to the mechanism of gene regulation - not by way of transcription factors binding to specific sequences - but rather, by factors being sterically blocked from binding by the 3-dimensional superstructure of the chromosome that occurs when histone proteins are deacetylated. The team suggests that as the brain ages, it becomes more difficult to recruit HDAC1,2 to the promoters needed to shut down the expression of the differentiation inhibitors. The results pose a confound for the certain applications of inhibitors of histone deacetylases (HDACi) which have demonstrated anti-tumor activity - but may - as suggested by this article - have negative consequences on brain repair processes. Image via Wikipedia Siming Shen et al., in their paper, "Age-dependent epigenetic control of differentiation inhibitors is critical for remyelination efficiency" provide insight on basic mechanisms of myelination. While myelination (think of it as the plastic insulation on copper electrical wires) makes normally developing neural networks much more efficient, it has a way of inhibiting the re-development and repair of mature neural circuits. The research team shows that recruitment of histone deacetylases (HDACs) is rather inefficient in mature oligodendrocytes precursor cells (the cells that adhere to bare neuronal axons and form the insulating myelin-rich sheath) in contrast to younger cells which differentiate readily. HDAC1 and HDAC2 are shown to down-regulate of Hes5 and Sox2, which have previously been implicated in blocking the differentiation of stem cells to oligodendrocytes. Here, the term 'epigenetic' refers to the mechanism of gene regulation - not by way of transcription factors binding to specific sequences - but rather, by factors being sterically blocked from binding by the 3-dimensional superstructure of the chromosome that occurs when histone proteins are deacetylated. The team suggests that as the brain ages, it becomes more difficult to recruit HDAC1,2 to the promoters needed to shut down the expression of the differentiation inhibitors. The results pose a confound for the certain applications of inhibitors of histone deacetylases (HDACi) which have demonstrated anti-tumor activity - but may - as suggested by this article - have negative consequences on brain repair processes. |
| Posted: 05 Sep 2008 06:18 PM CDT  Image via Wikipedia The mitogenic activities of the vascular endothelial growth factor protein family are well researched. A number of findings have linked this gene to learning and memory and hippocampal-dependent response to antidepressant medication. Indeed, its reasonable to expect that a mitogen such as VEGF would regulate hippocampal cell division and the accompanying benefits of new brain cells. Using high resolution structural MRI, Blumberg and team report evidence for such in their paper, "Influence of Vascular Endothelial Growth Factor Variation on Human Hippocampus Morphology". Individuals with the CC genotype at rs833070 and rs2146323 - located in the intron of the VEGF-A gene displayed smaller hippocampal volumes than T-allele and A-allele carriers, respectively. These 2 snps lie in a haplotype block with rs833068 which was assayed in my 23andMe profile - indicating that I happen to carry the TT genotype at rs833070 giving me slightly larger, more neurogenic & resilient hippocampus - I suppose. Now, if I could just figure out a way to put it to good use ! Image via Wikipedia The mitogenic activities of the vascular endothelial growth factor protein family are well researched. A number of findings have linked this gene to learning and memory and hippocampal-dependent response to antidepressant medication. Indeed, its reasonable to expect that a mitogen such as VEGF would regulate hippocampal cell division and the accompanying benefits of new brain cells. Using high resolution structural MRI, Blumberg and team report evidence for such in their paper, "Influence of Vascular Endothelial Growth Factor Variation on Human Hippocampus Morphology". Individuals with the CC genotype at rs833070 and rs2146323 - located in the intron of the VEGF-A gene displayed smaller hippocampal volumes than T-allele and A-allele carriers, respectively. These 2 snps lie in a haplotype block with rs833068 which was assayed in my 23andMe profile - indicating that I happen to carry the TT genotype at rs833070 giving me slightly larger, more neurogenic & resilient hippocampus - I suppose. Now, if I could just figure out a way to put it to good use ! |
| Posted: 05 Sep 2008 06:16 PM CDT  Image via Wikipedia It has been reported that cigarettes can impart some calm and clarity from racing thoughts and mental fog. Patients with schizophrenia, who often experience cognitive disorganization, are 2-4 times more likely than the general population to smoke, and also seem to prefer stronger brands of cigarettes. This is not surprising since nicotine can raise levels of dopamine indirectly via stimulation of alpha4/beta2 high affinity nicotinic acetyl choline receptors (nAChR) expressed widely in the parietal cortex of the human brain. In an open access article entitled, "Association of attentional network function with exon 5 variations of the CHRNA4 gene", Georg Winterer and colleagues demostrate that individuals who vary in a synonymous G/A variant (rs1044396) in the CHRNA4 gene - an snp which has previously been associated with nicotine dependence - show differential brain activity in the parietal cortex. When asked to remain alert and respond to rare visual "oddball" stimuli (visual oddball detection task), subjects with the AA genotype showed robust brain activity in the parietal cortex while subjects with the GG genotype showed very little change in activity. This finding reveals where in the brain - circuits connecting to the parietal cortex - may be especially important in mediating self-medication and even in the management of side-effects in psychiatric pharmacotherapy. Although rs1044396 is not measured in my 23andMe profile, the neighboring rs3787138 showing tight LD is measured and reveals that I am a boring, middle of the road heterozygote. As such, I do admit that I could use some mind-clearing relief from time to time - but, the yellow teeth are not quite worth it. Image via Wikipedia It has been reported that cigarettes can impart some calm and clarity from racing thoughts and mental fog. Patients with schizophrenia, who often experience cognitive disorganization, are 2-4 times more likely than the general population to smoke, and also seem to prefer stronger brands of cigarettes. This is not surprising since nicotine can raise levels of dopamine indirectly via stimulation of alpha4/beta2 high affinity nicotinic acetyl choline receptors (nAChR) expressed widely in the parietal cortex of the human brain. In an open access article entitled, "Association of attentional network function with exon 5 variations of the CHRNA4 gene", Georg Winterer and colleagues demostrate that individuals who vary in a synonymous G/A variant (rs1044396) in the CHRNA4 gene - an snp which has previously been associated with nicotine dependence - show differential brain activity in the parietal cortex. When asked to remain alert and respond to rare visual "oddball" stimuli (visual oddball detection task), subjects with the AA genotype showed robust brain activity in the parietal cortex while subjects with the GG genotype showed very little change in activity. This finding reveals where in the brain - circuits connecting to the parietal cortex - may be especially important in mediating self-medication and even in the management of side-effects in psychiatric pharmacotherapy. Although rs1044396 is not measured in my 23andMe profile, the neighboring rs3787138 showing tight LD is measured and reveals that I am a boring, middle of the road heterozygote. As such, I do admit that I could use some mind-clearing relief from time to time - but, the yellow teeth are not quite worth it. |
| Economist predicts "costectomies" as healthcare goes global [biomarker-driven mental health 2.0] Posted: 05 Sep 2008 06:14 PM CDT Just a pointer to a great overview article of trends in medical tourism at Economist magazine. Hope the price of jet fuel doesn't put a damper on these exciting trends. |
| Posted: 05 Sep 2008 06:11 PM CDT  Image via Wikipedia Nowadays, as many folks peer into the vast tangled thicket of their own genetic code, they, as I, assuredly wonder what it all means and how best to ascertain their health risks. One core theme that emerges from repeated forays into one's own data is that many of us carry a scads of genetic risk for illness, but somehow, find ourselves living rather normal, healthy lives. How can this be ? A recent example of this entails a C/T snp (rs35753505) located in the 5' flanking region of the neuregulin 1 gene which has been repeatedly associated with schizophrenia. Axel Krug and colleagues recently reported in their paper, "Genetic variation in the schizophrenia-risk gene neuregulin1 correlates with differences in frontal brain activation in a working memory task in healthy individuals" that T/C variation at this snp is associated with activation of the frontal cortex in healthy individuals. Participants were asked to keep track of a series of events and respond to a particular event that happened "2 events ago" . These so-called n-back tasks are not easy for healthy folks, and demand a lot of mental focus - a neural process that depends heavily on circuits in the frontal cortex. Generally speaking, as the task becomes harder, more activity in the frontal cortex is needed to keep up. In this case, individuals with the TT genotype seemed to perform the task while using somewhat less activity in the frontal cortex, rather than the risk-bearing CC carriers. As someone who has tried and failed to succeed at these tasks many times before, I was sure I would be a CC, but the 23andMe data show me to be a non-risk carrying TT. Hmmm ... maybe my frontal cortex is just underactive. Image via Wikipedia Nowadays, as many folks peer into the vast tangled thicket of their own genetic code, they, as I, assuredly wonder what it all means and how best to ascertain their health risks. One core theme that emerges from repeated forays into one's own data is that many of us carry a scads of genetic risk for illness, but somehow, find ourselves living rather normal, healthy lives. How can this be ? A recent example of this entails a C/T snp (rs35753505) located in the 5' flanking region of the neuregulin 1 gene which has been repeatedly associated with schizophrenia. Axel Krug and colleagues recently reported in their paper, "Genetic variation in the schizophrenia-risk gene neuregulin1 correlates with differences in frontal brain activation in a working memory task in healthy individuals" that T/C variation at this snp is associated with activation of the frontal cortex in healthy individuals. Participants were asked to keep track of a series of events and respond to a particular event that happened "2 events ago" . These so-called n-back tasks are not easy for healthy folks, and demand a lot of mental focus - a neural process that depends heavily on circuits in the frontal cortex. Generally speaking, as the task becomes harder, more activity in the frontal cortex is needed to keep up. In this case, individuals with the TT genotype seemed to perform the task while using somewhat less activity in the frontal cortex, rather than the risk-bearing CC carriers. As someone who has tried and failed to succeed at these tasks many times before, I was sure I would be a CC, but the 23andMe data show me to be a non-risk carrying TT. Hmmm ... maybe my frontal cortex is just underactive. |
| Benevolent dictators have longer AVPR1a promoters [biomarker-driven mental health 2.0] Posted: 05 Sep 2008 06:10 PM CDT The small neuropeptides oxytocin (OT) and arginine-vasopressin (AVP) are well known for their influence on promoting warm-and-fuzzy social behaviors in mammals. The G-protein coupled OTR and AVPR1a receptors are also the subject of much research in this area - particularly AVPR1a - since it shows differences in brain expression in polygamous vs. monogamous vole species, and also shows genetic associations with dysfunction in human social affiliation. In a recent foray into this line of research, Richard Ebstein and colleagues examine whether an individual's willingness to give away a cache of money is related to genetic variation in the promoter of the AVPR1a. In their paper, "Individual differences in allocation of funds in the dictator game associated with length of the arginine vasopressin 1a receptor RS3 promoter region and correlation between RS3 length and hippocampal mRNA", the researchers asked 203 college students to play the "dictator game" where, simply, one person gets a sum of money and can choose to keep it or give some of it away to the other player. Thats it. Give some of it away if you like, or just walk away with all of it, no questions asked & no consequences (your identity and the identity of the other player are masked). Amazingly, individuals actually DO give some of the money away (15% gave none of it away, 35% gave half away and 7% gave all - yes, all of it - away) ... and more amazing still ... those with longer stretches of microsatellite repeats at the RS1 & RS3 promoter sites in the AVPR1a gene, gave away significantly more money than individuals with shorter version of the repeats. |
| Posted: 05 Sep 2008 06:07 PM CDT  Image via Wikipedia Like most parents, I enjoy watching my children develop and marvel at the many similarities they bear to myself and my wife. The reshuffling of physical and behavioral features is always a topic of discussion and is the definitive icebreaker during uncomfortable silences with the inlaws. In some cases, the children are blessed with the better traits, but in other cases, there's no option but to cringe when, "Look - wow, he really has your nose - hmmm", is muttered. Most interesting, is the unfolding of patterns of behavior that unfold slowly with age. Differences in temperament and personality can instill great pride in parents but also can be a grating source of friction. One of my F1's has recently taken to sessions of shrill, spine rattling, screaming which I hope will pass soon. Image via Wikipedia Like most parents, I enjoy watching my children develop and marvel at the many similarities they bear to myself and my wife. The reshuffling of physical and behavioral features is always a topic of discussion and is the definitive icebreaker during uncomfortable silences with the inlaws. In some cases, the children are blessed with the better traits, but in other cases, there's no option but to cringe when, "Look - wow, he really has your nose - hmmm", is muttered. Most interesting, is the unfolding of patterns of behavior that unfold slowly with age. Differences in temperament and personality can instill great pride in parents but also can be a grating source of friction. One of my F1's has recently taken to sessions of shrill, spine rattling, screaming which I hope will pass soon.Why ? Many parents ask. "Have WE been raising him/her to do this ? - surely that's what the neighbors must think". "Is it something in the family ? I heard Aunt Marie was a bit of a screamer as a child - hmmm." In one of several of their landmark studies on the genetic regulation of pediatric brain development, Jay Giedd and colleagues, now provide in their recent paper, "Variance Decomposition of MRI-Based Covariance Maps Using Genetically-Informative Samples and Structural Equation Modeling", a core framework on the relative contribution of genes vs. environment for the developing cortex. The paper is part of an ongoing longitudinal study of pediatric brain development at the Child Psychiatry branch at NIMH. Some 600 children participated - including identical twins, fraternal twins, siblings and singleton children. The team used an analytical approach known as MACAAC (Mapping Anatomical Correlations Across the Cerebral Cortex) to ask how much does the variation in a single part of the brain co-vary with other parts ? Then the team used structural equation modeling to explore how much this co-variation might differ across identical twins vs. fraternal vs. siblings vs. age-matched singleton children. In locations where there is an high genetic contribution to co-variation in cortical thickness, identical twins should co-vary more tightly than fraternal twins or siblings etc. In this way, the team were able to parse out the relative influence of genes vs. environment to the developing brain. In general terms, the team reports that a single genetic factor accounts for the majority of variation in cortical thickness, which they note may be consistent with a major mechanism of development of cortical layers involving the migration of neurons along radial glia. Genetic co-variances across separate locations in the brain were highest in the frontal cortex, middle temporal gyrus and left supramarginal gyrus. Interestingly, when environmental covariations were observed, they were usually restricted to just one hemisphere, while genetic covariations were often observed bilaterally. Figure 5 of this paper is really incredible, it shows which areas of the cortex are more influenced by genes vs. environment. If I can just find the areas involved in screaming, the next time one of my neighbors looks askance at my F1, I'll be able to explain. |
| Posted: 05 Sep 2008 06:06 PM CDT  Image via Wikipedia Indeed, learning how to manage one's response to the negative emotions of others and stay out of trouble is an important life skill. At some point, most of us learn to just avoid angry, mean or melodramatically negative people and save ourselves the strife. Roy Perlis and colleagues, in their recent paper, "Association of a Polymorphism Near CREB1 With Differential Aversion Processing in the Insula of Healthy Participants", show how the transcriptional regulator CREB might exert an influence on this learning process. By having subjects view images of various facial expressions, the investigators found that individuals with the TT genotype at rs4675690 (C/T) showed less negative activation in the left insula, a brain region that is known to activate when subjects feel disgust, but not happiness, desire or fear. Subjects with the TT genotype have been shown to require more effort in the management of negative emotions and are at greater risk for suicide when being treated for depression. In the Perlis et al., study, TT subjects showed less of an effort (as measured in key presses) to avoid viewing emotionally distressing pictures. The known role of CREB in neural plasticity suggests that this gene may facilitate neural changes associated with memory. Unfortunately, 23andMe does not cover this SNP, so I'll just have to hope that (during the upcoming election) my insula keeps me on the path to enlightenment. Image via Wikipedia Indeed, learning how to manage one's response to the negative emotions of others and stay out of trouble is an important life skill. At some point, most of us learn to just avoid angry, mean or melodramatically negative people and save ourselves the strife. Roy Perlis and colleagues, in their recent paper, "Association of a Polymorphism Near CREB1 With Differential Aversion Processing in the Insula of Healthy Participants", show how the transcriptional regulator CREB might exert an influence on this learning process. By having subjects view images of various facial expressions, the investigators found that individuals with the TT genotype at rs4675690 (C/T) showed less negative activation in the left insula, a brain region that is known to activate when subjects feel disgust, but not happiness, desire or fear. Subjects with the TT genotype have been shown to require more effort in the management of negative emotions and are at greater risk for suicide when being treated for depression. In the Perlis et al., study, TT subjects showed less of an effort (as measured in key presses) to avoid viewing emotionally distressing pictures. The known role of CREB in neural plasticity suggests that this gene may facilitate neural changes associated with memory. Unfortunately, 23andMe does not cover this SNP, so I'll just have to hope that (during the upcoming election) my insula keeps me on the path to enlightenment.Update: Thanks so very much Brian for the info on rs7591784. This explains a lot - I'm a GG here, which means I'm a TT at rs4675690 - and have always had difficulty handling it when folks are rude to me. |
| Posted: 05 Sep 2008 06:03 PM CDT  Image via Wikipedia The recent SNP association report, "Identification of loci associated with schizophrenia by genomewide association and follow-up" (doi:10.1038/ng.201) by O'Donovan et. al, - an analysis of more than 370,000 Affymetrix SNPs on a population of 479 affected individuals - finds strong evidence for rs1344706 in the zinc finger protein 804A (ZNF804A). One clue to the otherwise inscrutable history of this gene may lie in the findings of a yeast 2-hybrid screen where ataxin-1 was used as a bait. Mutations in ATXN1 can give rise to Spinocerebellar Ataxia, a degenerative condition of the cerebellum and spinal cord. Such profound developmental deficits, even if weakly expressed would be consistent with the many cognitive difficulties experienced by patients with schizophrenia. Image via Wikipedia The recent SNP association report, "Identification of loci associated with schizophrenia by genomewide association and follow-up" (doi:10.1038/ng.201) by O'Donovan et. al, - an analysis of more than 370,000 Affymetrix SNPs on a population of 479 affected individuals - finds strong evidence for rs1344706 in the zinc finger protein 804A (ZNF804A). One clue to the otherwise inscrutable history of this gene may lie in the findings of a yeast 2-hybrid screen where ataxin-1 was used as a bait. Mutations in ATXN1 can give rise to Spinocerebellar Ataxia, a degenerative condition of the cerebellum and spinal cord. Such profound developmental deficits, even if weakly expressed would be consistent with the many cognitive difficulties experienced by patients with schizophrenia. |
| Posted: 05 Sep 2008 06:00 PM CDT A pair of Nature papers (here and here) find that mapping the risk of schizophrenia to the genome is more readily achieved when examining structural variation (insertions, deletions, duplications etc.). This is welcome news given the sparse success of SNP screening, although it would be reasonable to assume that SNPs can modify such structural variants (here for the most recent schizophrenia SNP association study). The pair of papers found similar sites, which is pretty amazing given that many structural variants are rare (see the 2006 survey report). The Copy Number Variation Project provides more details on this important class of variation. |
| Posted: 05 Sep 2008 05:57 PM CDT  Image by wallyg via Flickr The Manhattan Institute policy think-tank posts some commentary (including one by yours truly) on their Medical Progress Today section pertaining to the recent regulatory steps (backward). Image by wallyg via Flickr The Manhattan Institute policy think-tank posts some commentary (including one by yours truly) on their Medical Progress Today section pertaining to the recent regulatory steps (backward). |
| Posted: 05 Sep 2008 05:53 PM CDT Just re-posting from Gizmodo ... this looks like a positive step ... a medical chumby. |
| Posted: 05 Sep 2008 05:51 PM CDT  Image via Wikipedia James Schummers, Hongbo Yu and Mriganka Sur present their measurements of Ca++ dynamics in response to visual stimuli in the awake ferret and reveal highly refined patterns of astrocyte activity in their paper, "Tuned Responses of Astrocytes and Their Influence on Hemodynamic Signals in the Visual Cortex" (DOI: 10.1126/science.1156120). The genetic regulation of neurovascular coupling is key to understanding the way in which genetic variation may regulate brain function (or at least function as measured by the BOLD response). A closer look at BOLD response and genetic pathways that mediate astrocyte function would be music to my ears - or at least my auditory cortex astocytes. Image via Wikipedia James Schummers, Hongbo Yu and Mriganka Sur present their measurements of Ca++ dynamics in response to visual stimuli in the awake ferret and reveal highly refined patterns of astrocyte activity in their paper, "Tuned Responses of Astrocytes and Their Influence on Hemodynamic Signals in the Visual Cortex" (DOI: 10.1126/science.1156120). The genetic regulation of neurovascular coupling is key to understanding the way in which genetic variation may regulate brain function (or at least function as measured by the BOLD response). A closer look at BOLD response and genetic pathways that mediate astrocyte function would be music to my ears - or at least my auditory cortex astocytes. |
| Posted: 05 Sep 2008 05:49 PM CDT  Image by talkradionews via Flickr Amidst his hectic schedule managing the ongoing credit crisis, the New York Times notes that Federal Reserve chairman Ben Bernanke opened a bipartisan symposium which will, "lay the groundwork for what leaders of both parties predict will be a major push for health care legislation next year." From the article, it seems that since healthcare accounts for such a large (and growing) slice of the federal budget pie, fixing inefficiency and disparity in healthcare will assuredly involve more legislation and regulation on a growing scale. The article bummed me out since I suppose it portends new and shifting regulatory layers that will just make it harder for entrepreneurial consumer-driven and health2.0 innovations to establish themselves. Coincidental bummer that this symposium was taking place on the day NY and CA were stifling a new genetic testing industry. Image by talkradionews via Flickr Amidst his hectic schedule managing the ongoing credit crisis, the New York Times notes that Federal Reserve chairman Ben Bernanke opened a bipartisan symposium which will, "lay the groundwork for what leaders of both parties predict will be a major push for health care legislation next year." From the article, it seems that since healthcare accounts for such a large (and growing) slice of the federal budget pie, fixing inefficiency and disparity in healthcare will assuredly involve more legislation and regulation on a growing scale. The article bummed me out since I suppose it portends new and shifting regulatory layers that will just make it harder for entrepreneurial consumer-driven and health2.0 innovations to establish themselves. Coincidental bummer that this symposium was taking place on the day NY and CA were stifling a new genetic testing industry. |
| My 23andMe profile: no genomic guide to sports cheating [biomarker-driven mental health 2.0] Posted: 05 Sep 2008 05:48 PM CDT Just checked for my status at UGT2B17 - locus of a 150kb deletion variant covered here in the Economist - to see whether I might have missed my chance to inject 'roids and claim international sports glory without getting busted by the World Anti-Doping Agency. Unfortunately, the 23andMe profile provides data for the flanking genes YTHDC1 and UGT2A3 on chromosome 4, but not this specific gene. Doesn't quite seem to validate my choice to couch-potato-dom though. |
| Posted: 05 Sep 2008 05:45 PM CDT  Image via Wikipedia Amidst all the genome-wide 'snp-ing' going on of late (my 23-and-me data should arrive in a couple of weeks), Walsh and colleagues provide an incredible trove of structural variation (deletions/insertions in the size range of more than 100kbp but less than 100Mbp) that is 3- to 4-fold enriched in patients with adult onset and childhood onset schizophrenia. Their paper, "Rare Structural Variants Disrupt Multiple Genes in Neurodevelopmental Pathways in Schizophrenia" (DOI 10.1126/science.1155174) uses a variety of genome hybriziation techniques to map novel variants and finds, amazingly, that many of the hits are within genes that function in common pathways of brain development and synaptic function. The authors admit that it is hard to ascribe a population risk value to any one of these variants, but the biochemical pathways suggest that the genes that were identified by this method deserve a great deal of attention in the basic and clinical genetic research community. Image via Wikipedia Amidst all the genome-wide 'snp-ing' going on of late (my 23-and-me data should arrive in a couple of weeks), Walsh and colleagues provide an incredible trove of structural variation (deletions/insertions in the size range of more than 100kbp but less than 100Mbp) that is 3- to 4-fold enriched in patients with adult onset and childhood onset schizophrenia. Their paper, "Rare Structural Variants Disrupt Multiple Genes in Neurodevelopmental Pathways in Schizophrenia" (DOI 10.1126/science.1155174) uses a variety of genome hybriziation techniques to map novel variants and finds, amazingly, that many of the hits are within genes that function in common pathways of brain development and synaptic function. The authors admit that it is hard to ascribe a population risk value to any one of these variants, but the biochemical pathways suggest that the genes that were identified by this method deserve a great deal of attention in the basic and clinical genetic research community. |
| Posted: 05 Sep 2008 05:43 PM CDT  Image via Wikipedia Attention Deficit Hyperactivity Disorder is one of the most widespread psychiatric diagnoses in children. Parents who are faced with the decision to medicate or not medicate their children may wonder if their child - given a bit more time - won't just "grow out of it", as many children seem to do. With this in mind, it would obviously be helpful to have biomarkers that could predict whether certain children are more likely to simply acquire better attentional control on their own, and those children that might not. In their paper, "Polymorphisms of the Dopamine D4 Receptor, Clinical Outcome, and Cortical Structure in Attention-Deficit/Hyperactivity Disorder" (Arch Gen Psychiatry Vol 64 (no. 8), Aug 2007) a veritable dream team of child developmental neuroscientists working across several medical institutions report on two such biomarkers. One biomarker is the thickness of the orbitofrontal cortex and posterior parietal cortex. MRI-based measurments of these parts of the brain (just about 5mm thick!) show that children who carry a diagnosis of ADHD have a thinner cortical sheet in these regions. Another biomarker is genetic variation in an intracytoplasmic loop of the G-protein coupled dopamine D4 receptor (DRD4). Children with ADHD are more likely to carry a longer 7-repeat version of this VNTR polymorphism than the more common 4-repeat. Interestingly, the research team found that healthy children who carry the 7-repeat genetic variant also have slightly thinner cortex in the orbitofrontal and posterior parietal cortex, suggesting that this genetic variant may influence the risk of ADHD by way of an effect on cortical development. Additionally, the research team found that the cortex of ADHD children who carry this 7-repeat genetic variant "catches up" from age 8 and eventually falls within the range of healthy children by age 15. Lastly, the team reports that ADHD children who carry the 7-repeat had better clinical outcomes (albeit, many of the ADHD children in this study were treated with medication). Nevertheless, it appears that some progress has been made in identifying biomarkers that might predict favorable developmental trajectories. Image via Wikipedia Attention Deficit Hyperactivity Disorder is one of the most widespread psychiatric diagnoses in children. Parents who are faced with the decision to medicate or not medicate their children may wonder if their child - given a bit more time - won't just "grow out of it", as many children seem to do. With this in mind, it would obviously be helpful to have biomarkers that could predict whether certain children are more likely to simply acquire better attentional control on their own, and those children that might not. In their paper, "Polymorphisms of the Dopamine D4 Receptor, Clinical Outcome, and Cortical Structure in Attention-Deficit/Hyperactivity Disorder" (Arch Gen Psychiatry Vol 64 (no. 8), Aug 2007) a veritable dream team of child developmental neuroscientists working across several medical institutions report on two such biomarkers. One biomarker is the thickness of the orbitofrontal cortex and posterior parietal cortex. MRI-based measurments of these parts of the brain (just about 5mm thick!) show that children who carry a diagnosis of ADHD have a thinner cortical sheet in these regions. Another biomarker is genetic variation in an intracytoplasmic loop of the G-protein coupled dopamine D4 receptor (DRD4). Children with ADHD are more likely to carry a longer 7-repeat version of this VNTR polymorphism than the more common 4-repeat. Interestingly, the research team found that healthy children who carry the 7-repeat genetic variant also have slightly thinner cortex in the orbitofrontal and posterior parietal cortex, suggesting that this genetic variant may influence the risk of ADHD by way of an effect on cortical development. Additionally, the research team found that the cortex of ADHD children who carry this 7-repeat genetic variant "catches up" from age 8 and eventually falls within the range of healthy children by age 15. Lastly, the team reports that ADHD children who carry the 7-repeat had better clinical outcomes (albeit, many of the ADHD children in this study were treated with medication). Nevertheless, it appears that some progress has been made in identifying biomarkers that might predict favorable developmental trajectories. |
| rs4570625 - this is a really cool snp - if you're a nerd [biomarker-driven mental health 2.0] Posted: 05 Sep 2008 05:40 PM CDT Every student can recall at least one stereotypical professor who - while brilliant - kept the students amused with nervous and socially inept behavior. Let's face it, if you're in academia, you're surrounded by these - uh, nerds - and, judging by the fact that you are reading (not to mention writing) this blog right now - probably one of them. So, its natural to ask whether there might be a causal connection between emotionality, on the one hand, and cognitive performance on the other. Research on the neuromodulator serotonin shows that it plays a key role in emotional states - in particular, anxiety. Might it exert effects on cognitive performance ? In their paper, "A functional variant of the tryptophan hydroxylase 2 gene impacts working memory: A genetic imaging study", (DOI: 10.1016/j.biopsycho.2007.12.002) Reuter and colleagues use a genetic variation a G to T snp (rs4570625) in the tryptophan hydroxylase 2 gene, a rate limiting biosynthetic isoenzyme for serotonin to evaluate its effect on a cognitive task. They ask subjects (who are laying in an MRI scanner) to perform a rather difficult cognitive task called the N-back task where the participant must maintain a running memory queue of a series of sequentially presented stimuli. Previous research shows that individuals with the GG genotype show higher scores on anxiety-related personality traits and so Reuter and team ask whether these folks activate more or less of their brain when performing the N-back working memory task. It turns out that the GG group showed clusters of activity in the frontal cortex that showed less activation than the TT group. The authors suggest that the GG group can perform the task using by recruiting less of their brains - hence suggesting that perhaps there just might be a genetic factor that accounts for a possible negative correlation between efficient cognitive performance and emotionality. |
| Genetic ski patrol up and down the inverted U [biomarker-driven mental health 2.0] Posted: 05 Sep 2008 05:38 PM CDT  Image via Wikipedia The selection and dosing of medication in psychiatry is far from scientific - even though a great deal of hard science goes into the preclinical design and clinical development. One reason, among many, has to do with the so-called 'inverted-U-shaped' relationship between the dose of a psychoactive compound and an individuals' performance. Some folks show dramatic improvement with a given dose (their system may be functioning down at the low side of the inverted U mountain and hence, and added boost from medication may send their system up in performance), while others may actually get worse (those who are already at the peak of the inverted U mountaintop). How can a psychiatrist know where the patient is on this curve - will the medication boost raise or topple their patient's functioning ? Some insight comes in the form of a genetic marker closely linked to the DRD2 gene, that as been shown to predict response to a dopaminergic drug. Image via Wikipedia The selection and dosing of medication in psychiatry is far from scientific - even though a great deal of hard science goes into the preclinical design and clinical development. One reason, among many, has to do with the so-called 'inverted-U-shaped' relationship between the dose of a psychoactive compound and an individuals' performance. Some folks show dramatic improvement with a given dose (their system may be functioning down at the low side of the inverted U mountain and hence, and added boost from medication may send their system up in performance), while others may actually get worse (those who are already at the peak of the inverted U mountaintop). How can a psychiatrist know where the patient is on this curve - will the medication boost raise or topple their patient's functioning ? Some insight comes in the form of a genetic marker closely linked to the DRD2 gene, that as been shown to predict response to a dopaminergic drug.Michael Cohen and colleagues, in their European Journal of Neuroscience paper (DOI: 10.1111/j.1460-9568.2007.05947.x) entitled, "Dopamine gene predicts the brain's response to dopaminergic drug" began with a polymorphism linked to the DRD2 gene wherein one allele (TaqA1+) is associated with fewer DRD2 receptors in the striatum (these folks should show improvement when given a DRD2 agonist) while folks with the alternate allele (TaqA1-) were predicted to show a falling off of their DRD2 function in response to additional DRD2 stimulation. The research team then asked participants to perform a cognitive task - a learning task where subjects use feedback to choose between a 'win' or 'not win' stimulus - that is well known to rely on proper functioning of DRD2-rich frontal and striatal brain regions. Typically, DRD2 agonists impair reversal learning and, as expected, subjects in the low DRD2 level TaqA1+ genetic group actually got "more" impaired - or perseverated longer on rewarding stimuli and required more trials to switch on the go and figure out which stimulus was the "win" stimulus. Similarly, when differences in brain activity between baseline and positive "you win" feedback was measured, subjects in the drug treated, TaqA1+ genetic group showed an increase in activity in the putamen and the medial orbitofrontal cortex while subjects in the TaqA1- showed decreases in brain actiity in these regions. The authors suggest that the TaqA1+ group generally has a somewhat blunted response to positive feedback (sore winners) but that the medication enhanced the frontal-striatal reaction to positive feedback. Functional connectivity analyses showed that connectivity between the frontal cortex and striatum was worsened by the DRD2 agonist in the TaqA1+ genetic group and improved in the TaqA1- group. Although the interpretations of these data are limited by the complexity of the systems, it seems clear that the TaqA1 genetic marker has provided a sort of index of baseline DRD2 function that can be useful in predicting an individual's relative location on the theoretical inverted-U-shaped curve. |
| Posted: 05 Sep 2008 05:35 PM CDT  Image via Wikipedia I was just browsing the recent paper "Natural selection has driven population differentiation in modern humans" by Barreiro and colleagues (doi:10.1038/ng.78) and noticed in their supplementary table that the autism risk factor CNTNAP2 (as blogged about earlier here) contains at least one non-synonymous or 5'-UTR SNP with a high Fst value. Yann Klimenidis has a great post on this paper. Image via Wikipedia I was just browsing the recent paper "Natural selection has driven population differentiation in modern humans" by Barreiro and colleagues (doi:10.1038/ng.78) and noticed in their supplementary table that the autism risk factor CNTNAP2 (as blogged about earlier here) contains at least one non-synonymous or 5'-UTR SNP with a high Fst value. Yann Klimenidis has a great post on this paper. |
| Biomarker-friendly healthcare marketplace unveiled [biomarker-driven mental health 2.0] Posted: 05 Sep 2008 05:34 PM CDT Dr. Scott Shreeve has a great post on the launch of "Carol" a new, open & transparent healthcare marketplace. With DNA Direct offering services there, its easy to see how biomarkers and biomarker-driven care can work within a consumer-driven business model. Exciting to see the future today !! |
| Posted: 05 Sep 2008 05:31 PM CDT The publication of positive genetic association results is always something to cheer about although most of us know that on a different day, with a different sample, the results could just as easily been flat. So when a meta-analytical paper appears that actually supports a previous finding, you have to savor the sweet joy of it. So it is that Marcus Munafo and colleagues under the leadership of Jonathan Flint - one of the best meta-analytical assessors in the biz - report that the "C" allele of rs1800955 in the dopamine d4 receptor (DRD4) gene (doi: 10.1016/j.biopsych.2007.04.006) survives an analysis of several dozen studies on genetics and personality. Although small, "The evidence from our final meta-analysis indicatest hat the true effect size of the C-521T polymorphism on novelty seeking and impulsivity traits, if genuine, may account for up to 3%o of phenotypic variance." I, perhaps due to my "C" allele am excited ! MUNAFO, M. (2008). Association of the Dopamine D4 Receptor (DRD4) Gene and Approach-Related Personality Traits: Meta-Analysis and New Data. Biological Psychiatry, 63(2), 197-206. DOI: 10.1016/j.biopsych.2007.04.006 |
| Relief of big pharma's antidepressant blues is as easy as ABC ? [biomarker-driven mental health 2.0] Posted: 05 Sep 2008 05:29 PM CDT 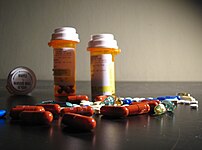 Image via Wikipedia Recent meta-analytical research, "Selective Publication of Antidepressant Trials and Its Influence on Apparent Efficacy" (N Engl J Med 2008;358:252-60) reveals that while 94% of published antidepressant drug trials show positive findings, only 51% of all such (published and unpublished) trials show positive effects (with a range of effect sizes from 11-69%). This is probably not surprising to patients and physicians (investors? ... well, maybe) who often search in vain, using trail and error, for a medication that can provide relief from major depression, one the the top disease burdens world-wide. Many have suggested that pharmacogenetics may provide a key to understanding the tremendous variability in medication response. For example, variations in the ABCB1, ATP-binding cassette sub-family B member 1, gene seem to predict who may show a response to certain antidepressants (citalopram, paroxetine, amitriptyline, and venlafaxine) medications, that are shuttled across the blood-brain-barrier endothelial membrane by ABCB1. In a pharmacogenetic medication trial involving 443 inpatients with depression who were treated at the Max Planck Institute of Psychiatry, the SNPs 2032583, rs2235015, rs2032583 and rs2235015 predict significantly different time course of response to treatment over 6 weeks. The paper, "Polymorphisms in the Drug Transporter Gene ABCB1 Predict Antidepressant Treatment Response in Depression" (doi: 10.1016/j.neuron.2007.11.017) is an example of pure and applied science at is best. The results pose a vexing dilemma for "really big" pharma however since the market size of genetic responders is obviously much smaller than market at large. Nevertheless, it seems inexorable change is underway. Image via Wikipedia Recent meta-analytical research, "Selective Publication of Antidepressant Trials and Its Influence on Apparent Efficacy" (N Engl J Med 2008;358:252-60) reveals that while 94% of published antidepressant drug trials show positive findings, only 51% of all such (published and unpublished) trials show positive effects (with a range of effect sizes from 11-69%). This is probably not surprising to patients and physicians (investors? ... well, maybe) who often search in vain, using trail and error, for a medication that can provide relief from major depression, one the the top disease burdens world-wide. Many have suggested that pharmacogenetics may provide a key to understanding the tremendous variability in medication response. For example, variations in the ABCB1, ATP-binding cassette sub-family B member 1, gene seem to predict who may show a response to certain antidepressants (citalopram, paroxetine, amitriptyline, and venlafaxine) medications, that are shuttled across the blood-brain-barrier endothelial membrane by ABCB1. In a pharmacogenetic medication trial involving 443 inpatients with depression who were treated at the Max Planck Institute of Psychiatry, the SNPs 2032583, rs2235015, rs2032583 and rs2235015 predict significantly different time course of response to treatment over 6 weeks. The paper, "Polymorphisms in the Drug Transporter Gene ABCB1 Predict Antidepressant Treatment Response in Depression" (doi: 10.1016/j.neuron.2007.11.017) is an example of pure and applied science at is best. The results pose a vexing dilemma for "really big" pharma however since the market size of genetic responders is obviously much smaller than market at large. Nevertheless, it seems inexorable change is underway. |
| Warfarin Response Testing: Medicare Calls for Feedback on Reimbursement [DNA Direct Talk] Posted: 05 Sep 2008 05:27 PM CDT Guest post from Trisha Brown, MS, CGC, DNA Direct’s VP of Clinical Affairs: FDA announced last year that the agency would update the label for the blood thinner warfarin to note that patients' genetic makeup could strongly influence their response to the drug. Too high a dose of warfarin, and patients may experience uncontrolled [...] |
| Stand Up To Cancer! [The Gene Sherpa: Personalized Medicine and You] Posted: 05 Sep 2008 05:23 PM CDT |
| "A must read for those interested in GMOs and/or the organic farming movement" [Tomorrow's Table] Posted: 05 Sep 2008 03:40 PM CDT Check out the review of Tomorrow's Table by evolutionary biologist Jonathan Eisen. Here are my favorite parts of his review (just a little cherry picking here): "I personally like the book a great deal, and enjoy how it switches back and forth between the authors (Pam and her husband Raoul Adamchak) and how it interweaves personal stories with discussion of the science and practice of organic farming and plant genetic engineering... ...the book really is a must read for those interested in GMOs and/or the organic farming movement as well those thinking about "slow food" and other related topics. In addition it is a wonderful personlized story, with a mixture of recipes, stories of research, discussions of teaching about organic agriculture, and some minor family drama. For the same reason that I like Amy Harmon's New York Times stories (such as the recent one on evolution) I like this book - it personalizes what is frequently a boring impersonal discussion..." |
| You are subscribed to email updates from The DNA Network To stop receiving these emails, you may unsubscribe now. | Email Delivery powered by FeedBurner |
| Inbox too full? | |
| If you prefer to unsubscribe via postal mail, write to: The DNA Network, c/o FeedBurner, 20 W Kinzie, 9th Floor, Chicago IL USA 60610 | |

![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=1bd0ba70-e9ae-4f8b-898c-3d017d8283f8)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=6f003976-3831-419f-a619-ef2dc04740bb)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=941d362d-c4b0-437c-a54d-e102f4a9bab0)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=42b7ace9-601f-4f07-91a9-14bfd423023e)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=2de532d7-8215-481e-8c4e-24468b7d6d1c)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=9f36e9b9-e00f-488f-a27f-a22024dd4054)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=b9e92f23-a157-4808-a773-52fae455f3e2)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=0e1a16b0-7616-49f0-9193-85b45c1610e9)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=a183a2c3-499a-4ac9-8d61-4b963ec02e35)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=986683e1-eddd-4186-a6ba-31a608793918)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=734d2d9f-cdf3-4d67-9222-677e01671c10)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=f998c118-4cf5-484c-902b-5db4dae86baf)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=d749c2da-69d5-41ab-9ccd-a07fc38c8eb0)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=c024b270-c0ce-4fac-8ca2-d97c8c430260)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=aac7da10-b62d-4115-83d3-9db5b7685851)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=169aca2d-b9f5-45bb-bac2-331f5bcc9f9c)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=a135ba21-7873-40c7-8636-7ff5100cd1e3)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=093965cc-b72b-4d55-9a90-e54bc3ed7fc2)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=8a1e8513-d5c9-49e8-aac3-9cc4e903fbfa)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=a9720d1f-db0f-445f-9ec7-fa57225b5339)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=c5c9e86e-0ef4-4186-a594-8b78fd8cbe29)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=629a2f6c-526c-4b8a-88f4-67b7eb216b78)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=62bb7f82-ee57-4285-90fb-c4fe2e118bee)
![Reblog this post [with Zemanta]](http://img.zemanta.com/reblog_e.png?x-id=d3ae944d-f489-4689-bc70-c85dee8dca8b)
1 comment:
Hi I'm a journalism student at New York University researching home genetic testing. I would love to interview you about your experience. Please email me at pdw222@nyu.edu.
Thank you!
Post a Comment